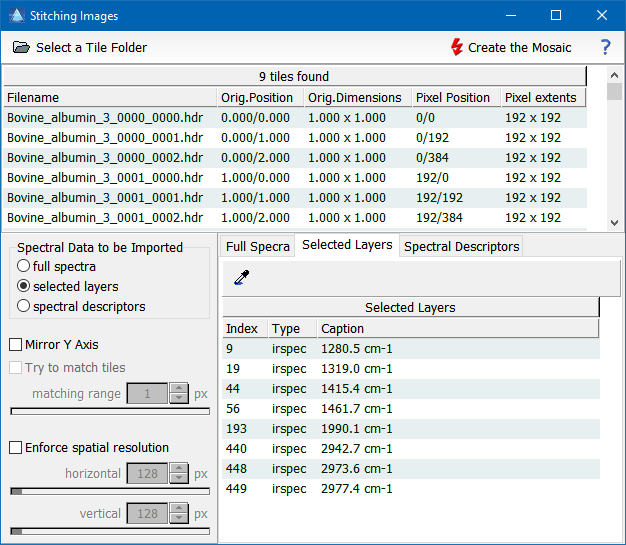
This dialog can be used to prepare any stitching operation. All the settings and selections made in the dialog are passed back to the calling program in the variable parameters of the function. The Mode parameter returns the stitching mode:
smFullSpec ... use all available layers
smSelLayers ... use only selected layers
smSpecDesc ... use spectral descriptors
The open array FNames returns the filenames of the selected tiles. The ImgCollection array contains the corresponding geometric parameters:
XPos ... position in x direction (in pixels)
YPos ... position in y direction (in pixels)
Width ... width of tile in pixels
Height ... height of tile in pixels
OrigPosX ... x position of the tile in "real world" coordinates
OrigPosY ... y position of the tile in "real world" coordinates
OrigWid ... the width of the tile in real coordinates
OrigHgt ... the height of the tile in real coordinates
The variable parameter HeaderFName returns the name of the header file which was used to generate the individual headers in the case AutoGenHDR was set to true. This header file is renamed to HeaderFName+'.$$$' after generating the individual headers.
The variable parameters EnforcedSpatialX and EnforcedSpatialY return the number of pixels along the x and the y axis if the checkbox "Enforce spatial resolution" has been checked, otherwise zero values are returned. Please note that this option has been implemented to correct a bug in the export interface of some IR microscopes which generate a wrong ENVI header under certain circumstances.
The open array LayersToExtract contains the selected layers (if the Mode is set to smSelLayers). All open array parameters are automatically resized to contain the information of all tiles.
The parameter AlignRange defines the range of pixels which the tiles are shifted to find an optimum fit. A zero value turns off the tile matching. The parameter Spdc contains the filename of the selected spectral descriptors (if Mode is smSpecDesc). The parameter MirrorY reflects the state of the checkbox "Mirror Y Axis".
The parameter CoordAnalysis contains the string extraction program which allows to defer the coordinates of the individual tiles from their file names. The extraction commands must comply with the function StringExtractor (which is called for the analyis) and must return two variables, OrigPosX and OrigPosY, containing the original positions of the upper left corner of the tiles. By default, StitchDialog expects the coordinates included in the file names as "[x,y]", i.e. aaaaaa[x,y]bbbbb, where aaaaaa and bbbbbb are valid strings not containing any square brackets and x and y are the coordinates (floating point or integer numbers). If the filenames exhibit this default structure, CoordAnalysis should be left empty, in any other case CoordAnalysis must contain code to extract x and y and emit it as variables OrigPosX and OrigPosY.
There are sometimes situations where a data source creates many datafiles (e.g. obtained from an FPA) but only a single (template) header file, which is valid for all tiles. In this case you can set the parameter AutoGenHDR to TRUE and load the file extension of the datafiles into DataFExt. This will automatically generate the corresponding header files from the template header file. In order to activate this option the folder containing the data files must be writable. The template file (XXXXX.HDR) is renamed by adding the file extension '.$$$' (i.e. the template file becomes XXXXX.HDR.$$$).
The parameter DebugFn specifies a debug file and should be left empty under normal circumstances. Setting it to a valid file path writes a protocol to this file which supports debugging of the string extraction.
The function returns the following error codes:
1 ... everything is OK, user pressed the "Stitch" button (1 = mrOK)
2 ... user closed the form without pressing the stitch button, parameters are undefined (2 = mrCancel)
-1 ... no sparse frequencies defined
-2 ... no descriptors defined
-3 ... no tiles selected
-4 ... AlignRange out of range (valid: 1..50)
-5 ... cannot autogenerate the header files
|

 Script Language
Script Language  I/O Functions
I/O Functions  Dialogs
Dialogs  StitchDialog
StitchDialog


 Script Language
Script Language  I/O Functions
I/O Functions  Dialogs
Dialogs  StitchDialog
StitchDialog
